
Research Activities
Research Activities
Home › Research Activities › Publications › iPS cells used to correct genetic mutations that cause muscular dystrophy
Publications
November 27, 2014
iPS cells used to correct genetic mutations that cause muscular dystrophy
Researchers at the Center for iPS Cell Research and Application (CiRA) show that induced pluripotent stem (iPS) cells can be used to correct genetic mutations that cause Duchenne muscular dystrophy (DMD) The research, published in Stem Cell Reports, demonstrates how engineered nucleases, such as TALEN and CRISPR, can be used to edit the genome of iPS cells generated from the skin cells of a DMD patient. The cells were then differentiated into skeletal muscles, in which the mutation responsible for DMD had disappeared.
DMD is a severe muscular degenerative disease caused by a loss-of-function mutation in the dystrophin gene. It inflicts 1 in 3500 boys and normally leads to death by early adulthood. Currently, very little is available in terms of treatment for patients outside palliative care. One option gaining interest is genomic editing by TALEN and CRISPR, which have quickly become invaluable tools in molecular biology. These enzymes allow scientists to cleave genes at specific locations and then modify the remnants to produce a genomic sequence to their liking. However, programmable nucleases are not pristine and often mistakenly edit similar sequences that vary a few base pairs from the target sequence, making them unreliable for clinical use because of the potential for undesired mutations.
For this reason, induced pluripotent stem cells (iPS cells) are ideal models, because they provide researchers an abundance of patient cells on which to test the programmable nucleases and find optimal conditions that minimize off-target modifications. CiRA scientists took advantage of this feature by generating iPS cells from a DMD patient. They used several different TALEN and CRISPR to modify the genome of the iPS cells, which were then differentiated into skeletal muscle cells. In all cases, dystrophin protein expression was convalesced, and in some cases, the dystrophin gene was fully corrected.
One key to the success was the development of a computational protocol that minimized the risk of off-target editing. The team built a database that all possible permutations of sequences up to 16 base pairs long. Among these, they extracted those that only appear once in the human genome, i.e. unique sequences. DMD can be caused by several different mutations; in the case of the patient used in this study, it was the result of the deletion of exon 44. The researchers therefore built a histogram of unique sequences that appeared in a genomic region that contained this exon. They found a stack of unique sequences in exon 45. According to Akitsu Hotta, who headed the project and holds joint positions at CiRA and the Institute for Integrated Cell-Materials Sciences at Kyoto University, "Nearly half the human genome consists of repeated sequences. So even if we found one unique sequence, a change of one or two base pairs may result in these other repeated sequences, which risks the TALEN or CRISPR editing an incorrect region. To avoid this problem, we sought a region that hit high in the histogram."
With this target, the team considered three strategies to modify the frame-shift mutation of the dystrophin gene: exon skipping by connecting exons 43 and 46 to restore the reading frame, frame shifting by incorporating insertion or deletion (indel) mutations, and exon knock-in by inserting exon 44 before exon 45. All three strategies effectively increased dystrophin synthesis in differentiated skeletal cells, but only the exon knock-in approach recovered the gene to its natural state. Importantly, editing showed very high specificity, suggesting that their computational approach can be used to minimize off-target editing by the programming nucleases.
Moreover, the paper provides a proof-of-principle for using iPS cell technology to treat DMD in combination with TALEN or CRISPR. The group now aims to expand this protocol to other diseases. First author Lisa Li explains, "We show that TALEN and CRISPR can be used to correct the mutation of the DMD gene. I want to apply the nucleases to correct mutations for other genetic-based diseases like point mutations."
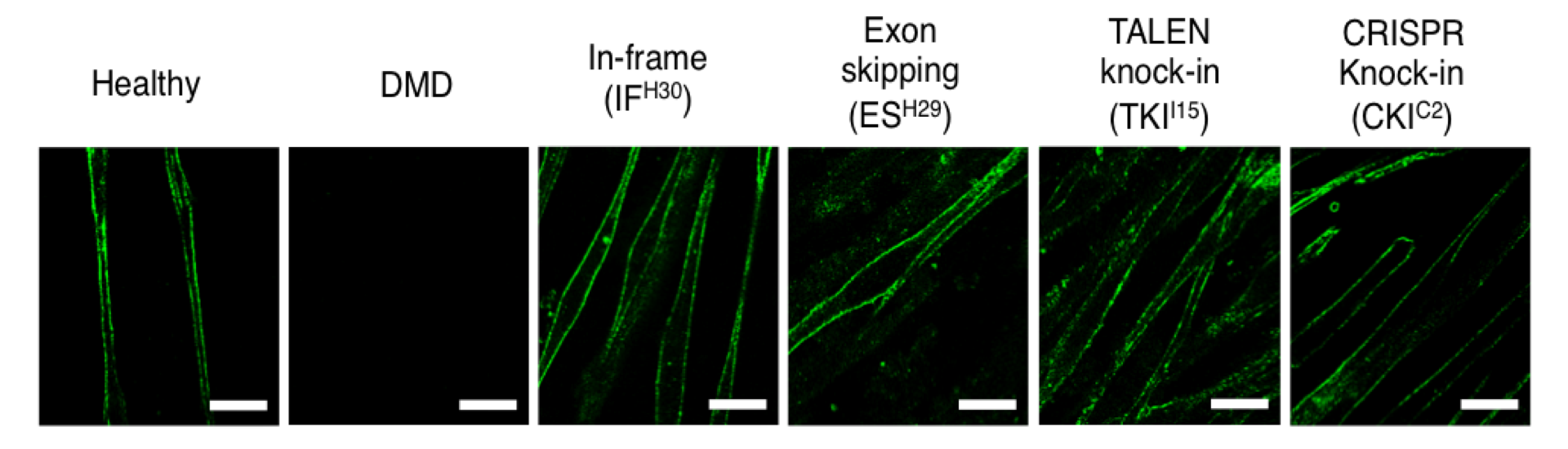
Figure
Immunofluorescence staining of skeletal cells differentiated from DMD-iPS cells. Untreated DMD skeletal cells do not express dystrophin (green) due to the deletion of exon 44. However, after any of the three correction strategies are applied to iPS cells, differentiation into skeletal cells results in normal dystrophin expression. Scale bar, 50 μm.
Details about the article
- Journal: Stem Cell Reports
- Title: Precise correction of the DYSTROPHIN gene in Duchenne Muscular Dystrophy patient-derived iPS cells by TALEN and CRISPR-Cas9
- Authors: Hongmei Lisa Li, Naoko Fujimoto, Noriko Sasakawa, Saya Shirai, Tokiko Ohkame, tetsushi Sakuma, Michihiro tanaka, Naoki Amano, Akira Watanabe, Hidetoshi Sakurai, Takashi Yamamoto, Shinya Yamanaka, and Akitsu Hotta






















