
News & Events
News & Events
News
January 29, 2018
A new technology to model disease mutations in cells
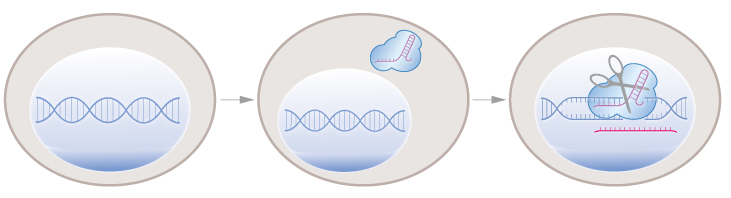
CRONUS uses two steps to control precisely genome editing at single base resolution. In the first step, Cas9 protein is expressed in the cell. In the second step, Cas9 protein is shuttled into the nucleus where the CRISPR-Cas9 system cuts the DNA.
Human DNA contains 3 billion base pairs, but a change in just one can have a profound effect. In fact, the majority of genetic diseases can be attributed to a single base pair mutation. Gene editing technology allows scientists to test the effects of these mutations, however, gene editing just a single base is technically difficult. A new study published in Scientific Reports describes CRONUS, gene-editing technology developed by researchers at CiRA that is expected to simplify the manufacturing of mutations, allowing scientists with rudimentary understanding of molecular biology to conduct gene-editing experiments for disease study.
CRONUS, or CRISPR-Cas9 regulated transcription and nuclear shuttling, takes advantage of the CRISPR-Cas9 system, a bacterial immune system that was first repurposed for genome editing less than six years ago. When CRISPR-Cas9 is activated, it cuts DNA in the cell at a specific region. Natural DNA repair machinery in the cell then fixes this damage. During the repair, scientists can manipulate the machinery to insert a desired mutation, thus permanently changing the DNA. There are two types of DNA repair machinery in human cells, homologous recombination (HR) and non-homologous end joining (NHEJ). Of the two, NHEJ repair is more common, but HR is preferred when studying gene function.
"NHEJ risks adding or cutting bases in the gene, which changes the protein size or in some cases prevents the protein from being expressed. HR gives specific mutations. If you are looking for new function of a gene, HR is preferred because you can dictate the type of mutations that are introduced," says Dr. Peter Gee, who co-authored the study.
CRONUS functions by using two drugs to regulate CRISPR-Cas9 activity.
"We designed CRONUS so that adding doxycycline expresses Cas9 protein and dexamethasone transports Cas9 into the nucleus," where it cuts the DNA, explains Junior Associate Professor Akitsu Hotta, who managed the project. The two-drug approach, added Hotta, significantly improves the accuracy of by controlling the Cas 9 activity to cleave DNA.
To demonstrate CRONUS, the Hotta lab inserted single-stranded oligodeoxynucleotides to the cells to create various mutations at the cut site. CRONUS was confirmed using multiple genes and multiple cell types, including iPS cells, with HR occurring in about one third of cases.
"In some systems you are lucky if you get over 3%. iCRISPR gives similar efficiency as CRONUS, but establishing the system takes a couple of months longer," said Gee.
The magnitude improvement in efficiency means one tenth the number of cells need to be cloned to conduct experiments, saving time and effort for the scientist.
Hotta said CRONUS should appeal to novice scientists for many reasons. "CRONUS is simple, affordable and saves time. We expect CRONUS to encourage more scientists to enter research on disease models caused by genetic variations."
Paper Details
- Journal: Scientific Reports
- Title: Site-specific randomization of a the endogenous genome by a regulatable CRISPR-Cas9 piggyBac system in human cells
- Authors: Kentaro Ishida1,2, Huaigeng Xu1, Noriko Sasakawa1,2, Mandy Siu Yu Lung1, Julia Alexandra Kudryashev3, Peter Gee1,4, and Akitsu Hotta1,2,4
- Author Affiliations:
- Center for iPS Cell Research and Application (CiRA), Kyoto University, Kyoto, Japan
- Core Center for iPS Cell Research, Research Center Network for Realization of Regenerative Medicine, Japan Agency for Medical Research and Development (AMED), Tokyo, Japan
- Massachusetts Institute of Technology, Cambridge, CA, USA
- Institute for integrated Cell-Material Sciences (iCeMS), Kyoto University, Kyoto, Japan






















