
News & Events
News & Events
News
March 13, 2020
A safer way to edit gene mutations
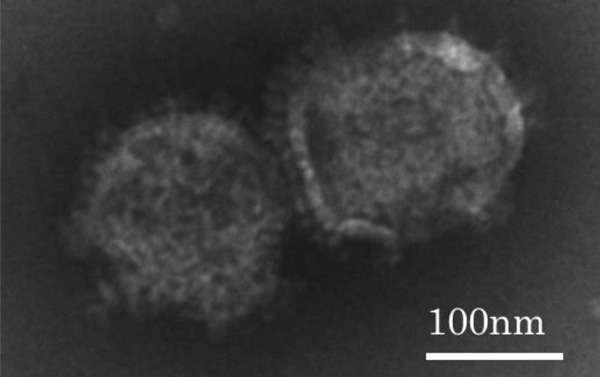
NanoMEDIC (130-140 nm)
CiRA scientists report a new molecular delivery system, NanoMEDIC (nanomembrane-derived extracellular vesicles for the delivery of macromolecular cargo), to insert genome editing protein tools into cells to correct gene mutations as a form of gene therapy. They test their system to repair cells harboring mutations that cause Duchenne muscular dystrophy (DMD), a disease in which muscles throughout the body progressively weaken.
All DMD is due to some type of mutation in the dystrophin gene, resulting in an absent dystrophin protein, which is essential for maintaining muscle integrity. One treatment recently approved is exon skipping, a technique that produces a functional but slightly truncated version of the dystrophin protein. Clinical trials for exon skipping using antisense oligonucleotides have been investigated, but this approach gives only a temporary effect and requires the patient to undergo multiple treatments (i.e. weekly injections). Exon skipping with gene editing, on the other hand, is permanent and in principle could remove the root-cause of the disease with just one treatment.
Genome editing has two intrinsic challenges before it can benefit patients, however. First is introducing the genome editing machinery into the target cell and second is to edit only the target gene. For the first problem, scientists have resorted to molecular tools found in viruses, since viruses have evolved to infect host cells effectively in nature.
To infect a cell, viruses bind to the cell surface, enter the cell as particles, and then pop inside, releasing their viral material like a reverse pimple. The CiRA scientists replaced the pathogenic viral material in the viral particles with molecules that can execute a desired effect, such as therapeutic genome editing.
The CRISPR-Cas9 system, currently the most popular of genome editing tools, consists of the Cas9 protein and sgRNA. Both these components are necessary for successful gene editing, which is why the CiRA scientists designed NanoMEDIC to encapsulate them into virus-like particles.
For the encapsulation of Cas9 protein, NanoMEDIC binds the protein to Gag, a protein that binds to the inside of the particles. Getting sgRNA into the same particle proved more difficult.
"Packaging the Cas9 protein went smoothly. We struggled to fill the particles with sgRNA into NanoMEDIC because we could not traffic the sgRNA from the nucleus to the plasma membrane," says Dr. Peter Gee, who was first author of the study.
Both Cas9 and sgRNA are made in the cell nucleus, but the particles are made at the plasma membrane. "It's a quite a journey from the nucleus to the membrane," says Gee.
Gag guided this journey for Cas9, because it binds to the plasma membrane region that forms the particles. However, to traffic the sgRNA was, Gee adds, "a lot of trial and error. We were able to design the sgRNA to contain a packaging signal targeting HIV Gag at the plasma membrane."
This packaging signal acted as a homing device to direct sgRNA into the same particle with Cas9. However, once encapsulating the two together, NanoMEDIC then needs to release sgRNA so that it can bind with Cas9 to activate the gene editing process before the particle purges its content into the target cell. The solution was to flank sgRNA with two ribozymes, which cut sgRNA free.
To test NanoMEDIC, the lab used skeletal muscle cells and iPS cells made from DMD patients along with other human cell types and also targeted several genes besides the dystrophin gene to confirm the versatility of the system. Indeed, the dystrophin protein could be recovered in the treated skeletal muscle cells and DMD patient's iPS cells.
Importantly, while the targeted gene was edited at rates comparable with other gene editing delivery methods, the rate of untargeted gene editing was much lower, suggesting a major increase in safety.
"The risk of cutting the non-target site with NanoMEDIC was almost undetectable, while maintaining the similar on-target activity with a conventional method. Since our NanoMEDIC disappears within cells quickly, this helps to reduce the adverse cutting risks, we believe." says CiRA Junior Associate Professor Akitsu Hotta, who led the study.
In fact, NanoMEDIC was cleared in mice within three days of injection. The quick clearance of NanoMEDIC should minimize the risk of residual CRISPR-Cas9 delivered from NanoMEDIC editing off-target genes. On the other hand, just one shot of NanoMEDIC maintained the gene-editing effect for more than five months. This long-lasting effect suggests the need for few if any additional treatments.
"We only measured up to five months, but the effect should be much longer," says Gee.
The key, he adds, is that NanoMEDIC provides the same quality effect on the target as current delivery systems but at far less risk.
"We want NanoMEDIC to become an attractive tool for delivering proteins into cells with minimal off-target risks."
Paper Details
- Journal: Nature Communications
- Title: Extracellular nanovesicles for packaging of CRISPR- Cas9 protein and sgRNA to induce therapeutic exon skipping
- Authors: Peter Gee1,2, Mandy S.Y. Lung1, Yuya Okuzaki1, Noriko Sasakawa1, Takahiro Iguchi1,
Yukimasa Makita3, Hiroyuki Hozumi3, Yasutomo Miura1, Lucy F. Yang1, Mio Iwasaki1, Xiou H. Wang1, Matthew A. Waller1, Nanako Shirai1, Yasuko O. Abe1, Yoko Fujita4, Kei Watanabe1, Akihiro Kagita1, Kumiko A. Iwabuchi1, Masahiko Yasuda4, Huaigeng Xu1, Takeshi Noda5, Jun Komano6,7,
Hidetoshi Sakurai1, Naoto Inukai3, and Akitsu Hotta1,2 - Author Affiliations:
- Center for iPS Cell Research and Application (CiRA), Kyoto University, 53 Kawahara-cho, Shogoin, Sakyo-ku, Kyoto 606-8507, Japan
- Institute for Integrated Cell-Material Sciences (iCeMS), Kyoto University, Yoshida Ushinomiya-cho, Sakyo-ku, Kyoto 606-8507, Japan
- T-CiRA Discovery, Takeda Pharmaceutical Company Limited, 26-1, Muraoka-Higashi 2-chome, Fujisawa, Kanagawa 251-8555, Japan
- Pathology Analysis Center, Central Institute for Experimental Animals, Kawasaki, Kanagawa 210-0821, Japan
- Laboratory of Ultrastructural Virology, Institute for Frontier Life and Medical Sciences, Kyoto University, 53 Kawahara-cho, Shogoin, Sakyo-ku, Kyoto 606-8507, Japan
- Department of Clinical Laboratory, Nagoya Medical Center, 1-1 4-chome, Sannomaru, Naka-ku, Nagoya, 460-0001, Japan
- Current address: Department of Infection Control, Osaka University of Pharmaceutical Sciences, 4-20-1 Nasahara, Takatsuki, Osaka, 569-1041, Japan






















