
News & Events
News & Events
―Reducing Variability through AI Prediction―
News
March 24, 2026
A New Method to Achieve Precise Gene Editing in Embryos
―Reducing Variability through AI Prediction―
Key Research Highlights
- Established a workflow to predict genome editing outcomes in embryos by combining AI (machine learning) models with preliminary validation in ES cells.
- Demonstrated high efficiency of desired genetic and phenotypic changes in first-generation (founder) mice following genome editing in embryos.
- Expected to shorten the production time of genome-edited animals, improve the reliability of results, and contribute to the reduction of laboratory animal use.
A research group—including graduate student Khanui Lkhagvadorj, Assistant Professor Eiichi Okamura (at the time of the research), and Professor Masatsugu Ema of the Animal Resource Center for Sciences at Shiga University of Medical Science; Professor Seiya Mizuno of the Laboratory Animal Resource Center at the University of Tsukuba; and Associate Professor Knut Woltjen of the Center for iPS Cell Research and Application at Kyoto University—has developed a new method to reduce the variability in editing outcomes following genome editing in embryos and more reliably introduce intended changes.
Conventional genome editing methods (such as CRISPR/Cas9※1) have faced difficulties in controlling repair outcomes after DNA is cut, requiring significant time and large numbers of animals to obtain individuals with the intended modifications.
In this study, the group established a procedure using the AI model "inDelphi※2" to predict probable changes in genome-edited embryos and validate those predictions first in cultured cells (ES cells※3). This workflow enabled the researchers to determine probable genetic changes in embryos in advance and identify the optimal genome-editing strategy to obtain the desired first-generation (founder) individuals※4 with high efficiency.
This method is expected to accelerate the production of genetically modified animals and enhance the accuracy of genome-editing results. Furthermore, the total number of animals used can be reduced by increasing the number of individuals with the desired genomic changes. For animals such as primates, which take considerably longer to obtain offspring, a highly efficient means of generating founder individuals with the desired genomic changes offers significant advantages.
These research results are scheduled to be published in the international scientific journal Communications Biology on March 23, 2026.
Genetically modified animals are indispensable for investigating the causes of human diseases and evaluating new treatments or drug candidates. In recent years, the use of genome editing technology to introduce genetic changes directly in embryos has become increasingly common, making it possible to produce experimental animals with disease-causing mutations found in human patients, for example, more rapidly than before.
Nonetheless, when genome editing is performed on embryos, the resulting changes can vary between individuals despite using the same design, or different changes may be mixed within a single individual (mosaicism※5), often failing to yield the intended results. In such cases, it is necessary to produce and examine many individuals to identify those meeting the research objectives, thereby increasing both the time required and the number of animals used.
Furthermore, a common approach to obtain experimental animals with identical genetic information is to produce offspring from founder individuals for detailed analyses. However, for animals with long generation times, such as primates, more time is required before research can even begin. Thus, to accelerate research, there has been a strong demand for innovations that enhance the success rate of obtaining the intended genetic modifications in founder individuals.
Against this background, establishing a method to perform embryo gene modification more efficiently, with lower variability in editing outcomes, and forecasting the resulting genomic changes to improve experimental efficiency has been a major challenge.
In this study, the team established a workflow to identify designs most likely to produce the intended change before performing gene modification in embryos.
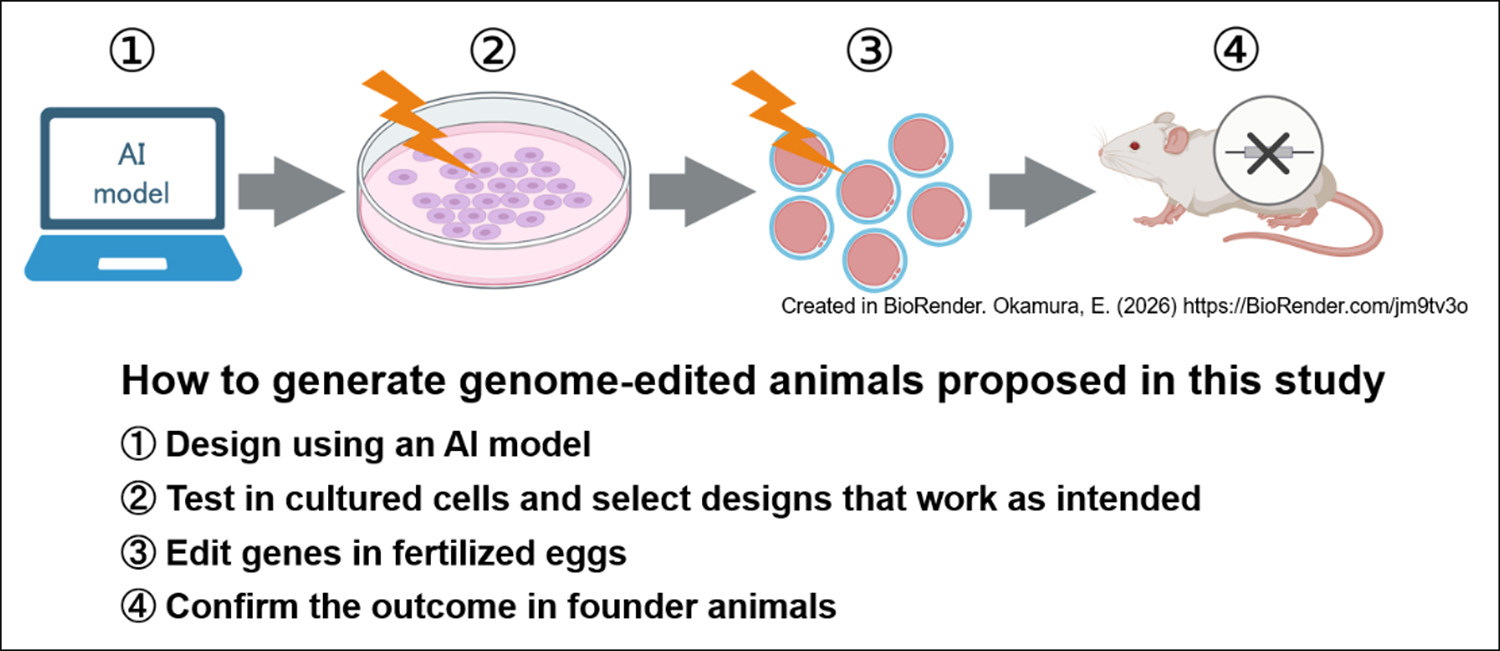
First, an AI model was used to predict probable post-modification changes by various design candidates for gene modification and select those most likely to lead to the intended change. Next, the shortlisted candidates were experimentally tested in cultured cells to confirm whether the predicted changes indeed occur. Finally, using designs deemed promising during this preliminary validation, gene modification was conducted in mouse embryos.
In experiments targeting multiple genes, the team demonstrated that they could successfully deactivate gene functions as intended (knockout※6) and observed the expected physical changes (phenotypes) in founder individuals with high efficiency.
Thus, this research moves embryonic gene modification one step beyond the "wait and see" process, presenting a practical procedure for selecting designs likely to succeed based on in silico prediction and in vitro confirmation, thereby hastening the experimental process.
This method is expected to speed up the workflow of producing genetically modified animals and increase confidence in obtaining the desired founder individuals. Additionally, because it can increase the number of individuals with the intended alterations, it will help limit the number of experimental animals required. Furthermore, research can progress more rapidly by first using ES cells as a proxy to confirm the resulting genome-editing changes before applying them to blastocysts or embryos. Particularly for animals like primates that require longer periods to generate offspring, a highly efficient embryonic genome-editing protocol dramatically reduces time and other experimental resources needed by reducing the need for additional breeding.
- Title
Optimizing CRISPR Precision in Mouse Embryos via Microhomology-Mediated End Joining-Dominant Targeting - Journal
Communications Biology - Authors
Khanui Lkhagvadorj#, Eiichi Okamura#,*, Taito Taki, Hayate Suzuki, Akihiro Kuno, Yasushi Itoh, Seiya Mizuno, Knut Woltjen*, Masatsugu Ema*
#: Co-First Authors
*: Corresponding Authors
This research was conducted with the support of the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Numbers: 25K02195, 24K18045) and the Japan Agency for Medical Research and Development (AMED) (Grant Number: JP223fa627008).
※1 CRISPR/Cas9
A genome editing technology that can modify specific genes by combining a "guide RNA (gRNA)" that recognizes a target DNA sequence with the "Cas9" enzyme that cuts the DNA. After cutting, the cell's repair mechanism works, resulting in changes such as deactivating (knocking out) the gene's function.
※2 inDelphi
A machine learning model reported in 2018 that predicts the likelihood of deletions and insertions (indels) resulting from cell repair after DNA is cut by CRISPR/Cas9, based on large-scale data (MV Shen, Nature, 2018).
※3 ES Cells (Embryonic Stem Cells)
Cells established from the blastocyst stage of an embryo that can differentiate into various types of cells. In this study, they were used as a validation system to determine the types of mutations most likely to occur with candidate gRNAs before performing editing in embryos.
※4 Founder Individuals
Individuals obtained after performing genome editing on an embryo. Typically, the next generation (F1) is produced to standardize the genotype, but if the intended mutation can be obtained in founder individuals, analysis can proceed more quickly.
※5 Mosaicism
A state in which cells with different genotypes coexist within the same individual. When genome editing is performed on an embryo, editing can occur at multiple stages of cell division during early development, possibly resulting in a mixture of different mutations within the individual and complicating analysis.
※6 Knockout
To deactivate the function of a gene. This state is created by cutting the gene with CRISPR/Cas9 and introducing mutations such as deletions during the repair process, ultimately leading to a loss of gene function.






















