
News & Events
News & Events
News
May 01, 2026
Uncovering a Translational Firewall That Maintains Adult Intestinal Stem Cell Identity
The intestinal epithelium is one of the body's most rapidly renewing tissues and depends on Lgr5-positive intestinal stem cells (ISCs) to continuously generate differentiated cell types that absorb nutrients and maintain barrier function. When this finely balanced system is disrupted, epithelial cells can revert to a fetal-like, regenerative state that helps repair acute injury but compromises normal tissue function if sustained. It remains unclear how adult stem cells normally resist this reprogramming.
In this study, the researchers focused on eIF4G2 (also known as DAP5 or NAT1), a non-canonical translation initiation factor known to regulate the translation of specific mRNAs but previously thought to play mainly supportive roles in protein synthesis. Using inducible mouse models and intestinal organoid systems, they found that deleting eIF4G2 in adult intestinal epithelium leads to a rapid collapse of Lgr5-positive ISCs and a failure of secretory cell maturation. Remarkably, overall tissue architecture—such as villus structure—was largely preserved, indicating that the epithelium shifted into an alternative, compensatory state rather than undergoing widespread damage.
Transcriptomic and single-nucleus multiomic analyses revealed that the loss of eIF4G2 drives intestinal cells into a stable fetal-like and regenerative program marked by strong activation of YAP-TEAD signaling. This state resembles developmental or injury-induced regeneration but, in contrast to normal repair processes, persists for months when eIF4G2 is absent. Importantly, this transition was not accompanied by classical inflammatory or stress-response pathways, underscoring that it is driven by an intrinsic regulatory mechanism.
At the molecular level, ribosome profiling showed that eIF4G2 loss does not reduce bulk protein synthesis. Instead, it selectively impairs translation efficiency of a specific group of mRNAs, particularly those encoding chromatin and epigenetic regulators. Among the most affected were CREBBP and EP300, the KAT3 family acetyltransferases responsible for key histone acetylation marks associated with active gene regulation. The reduced translation of these factors led to decreased histone acetylation and highly selective remodeling of enhancer landscapes: While adult ISC and WNT-Notch regulatory elements were decommissioned, fetal and YAP-associated enhancers became more active.
Pharmacological inhibition of CREBBP/EP300 closely mimicked the effects of eIF4G2 loss, reinforcing the idea that dampened epigenetic coactivator output is a central driver of the observed stem cell identity switch. Notably, fetal intestinal spheroids tolerated similar biochemical defects without loss of viability, highlighting a developmental stage-specific reliance on this translational buffering system.
Together, these findings identify eIF4G2 as a key molecular link between selective translation and epigenetic control that stabilizes adult intestinal stem cell identity. The work reveals that translation initiation factors can function as precision regulators of cell fate, rather than mere housekeeping components, and provides new insight into how adult tissues balance regeneration with long-term functional integrity.
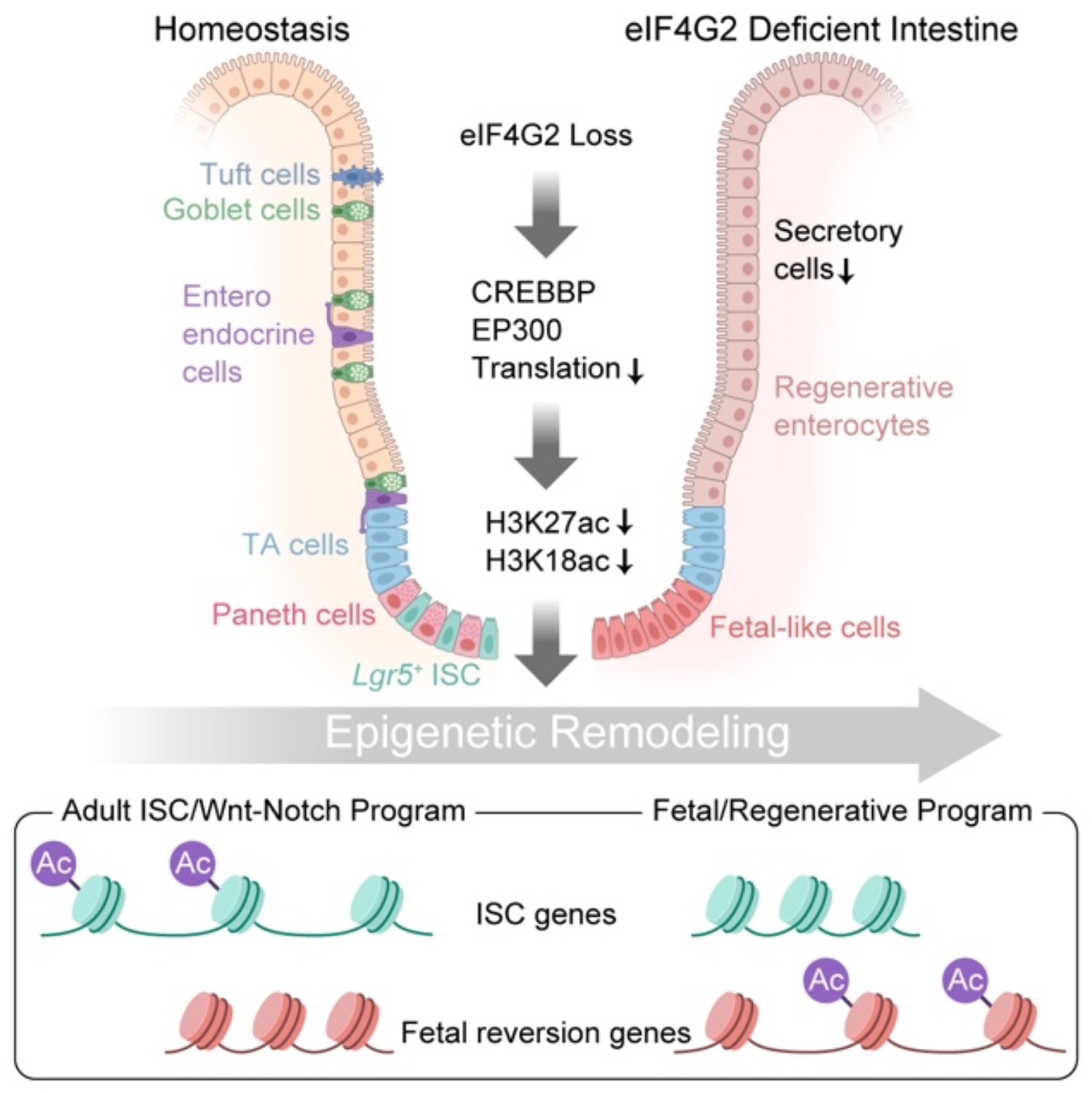
Graphical abstract
Paper Details
- Journal: Cell Stem Cell
- Title: eIF4G2-Mediated Selective Translation of Chromatin Regulators Safeguards Adult Intestinal Stem Cell Identity and Differentiation
- Authors:
Haruko Kunitomi1*, Aye Myat Khaine1, Radia Jamee1, Vanessa Arreola1, Mariselle Lancero1, Amba Raychaudhuri1, Samuel Perli1, Yoshiko Sato2, Mio Iwasaki2, Pedro Ruivo3, Kiichiro Tomoda1,2, Mari Mito4, Yuichi Shichino4,5, Shintaro Iwasaki4,6, Shinya Yamanaka1,2*
*: Corresponding authors - Author Affiliations:
- Gladstone Institute of Cardiovascular Disease, Gladstone Institutes
- Center for iPS Cell Research and Application, Kyoto University
- Comparative Pathology Laboratory, University of California, Davis
- RNA Systems Biochemistry Laboratory, Pioneering Research Institute, RIKEN
- Department of RNA Biochemistry, Institute of Medicine, University of Tsukuba
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences,
The University of Tokyo






















