
Research Activities
Research Activities
Publications
January 29, 2021
Flashing light on artificial RNA
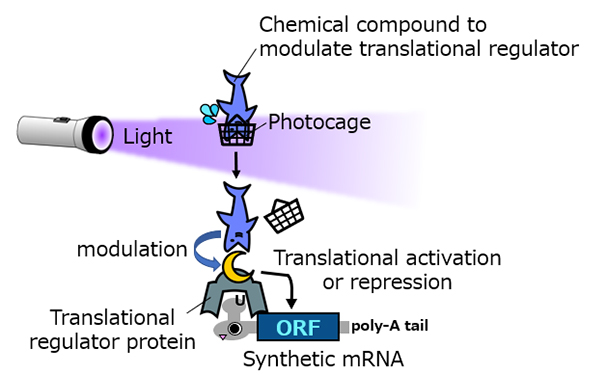
In the Central Dogma, while DNA is the blueprint of our bodies, its information is converted to proteins, which regulate the function of a cell. RNA is the key intermediary of this conversion. Biologists have realized that synthetic RNA transfected into a cell can be exploited to produce desired proteins that control how a cell behaves. However, turning this synthetic RNA on and off is equally crucial to get the desired effect. A new study by the
Hirohide Saito group reports new biotechnology known as Caliciviral VPg-based Translational Activator, or CaVT, that can be activated to translate target RNA into proteins.
The Saito lab actually reported CaVT earlier last year, but the activation depended on the use of small molecules. When imagining wider applications, Saito explains light is preferred.
"When we think of patient treatments, light has high spatial and temporal accuracy to target the sick cells while minimizing side effects," he said.
CaVT consist of two components. One, an RNA-binding protein, MS2CP, binds directly to the target RNA. The other, feline caliciviral VPg protein, is crucial for initiating the RNA translation.
In original CaVT, MS2CP and VPg are directly fused. To make the system sensitive to light, the researchers made two new versions. In one, they effectively cut CaVT in half, rendering it functionless. However, the function could be recovered by fusing a HaloTag protein to the VPg portion and an eDHFR protein to the MS2CP portion. When HaloTag and eDHFR interact, the CaVT is activated.
"HaloTag and eDHFR can bind to one another through a fused complex of trimethoprim and HaloTag ligand. Then the CaVT recovers to translate the protein," explained Dr. Hideyuki Nakanishi, an assistant professor at the Tokyo Medical and Dental University. He is also an inventor of CaVT and led the new study while working at CiRA.
To make this split CaVT sensitive to light, he protected the fused complex with a photocage. Photocages are molecules that respond to light. In biology, they are frequently used to trap certain molecules. Exposing the photocage to light releases the trapped molecule, i.e. the fused trimethoprim and HaloTag ligand. That allowed HaloTag and eDHFR to interact and CaVT to function.
In the second approach, Nakanishi attached a destabilizing domain to CaVT. As the name suggests, destabilizing domains destabilize a protein. In this design, it was selected to have an affinity for trimethoprim, which would prevent the destabilization. Again, by using photocaged trimethoprim, CaVT could be activated if the cell was exposed to UV light, which would release the trimethoprim, allowing it to bind to the destabilizing domain.
To test their light-sensitive CaVT, the researchers designed modified RNA that translate fluorescent proteins and transfected them into human cells. Without UV light, the cells remained relatively normal. However, once the cells were exposed to UV light, both CaVT translated the modified RNA, turning the cells green.
Functionally, the split CaVT and destabilizing domain-fused CaVT differed in two important ways. First, split CaVT could only activate translation, whereas destabilizing domain-fused CaVT could activate and repress translation depending on the structure of the modified RNA. Second, split CaVT was more robust to shortened ligand exposure time.
While in this preliminary study the modified RNAs were designed to translate fluorescent proteins, other proteins with physiological effects, such as causing the death of cancer cells, are equally feasible.
Nakanishi stresses that combining light and synthetic RNA could make for effective biotechnologies to manipulate cell behaviour.
"There is lots of research on synthetic RNA to control cell fate and function. RNAs are safe and easy to synthesize, but activating them is a challenge. Optically sensitive RNAs allow for precise spatiotemporal regulation," he said.
Paper Details
- Journal: Cell Chemical Biology
- Title: Light-controllable RNA-protein devices for translational regulation of synthetic mRNAs in mammalian cells
- Authors: Hideyuki Nakanishi1,2, Tatsuyuki Yoshii3,4, Shunsuke Kawasaki1, Karin Hayashi1, Keita Tsutsui3, Choji Oki5, Shinya Tsukiji3,5, Hirohide Saito1,6,*
- Author Affiliations:
- Department of Life Science Frontiers, Center for iPS Cell Research and Application, Kyoto University, 53 Kawahara-cho, Shogoin, Sakyo-ku, Kyoto 606-8507, Japan
- Department of Biofunction Research, Institute of Biomaterials and Bioengineering, Tokyo Medical and Dental University (TMDU), 2-3-10 Kanda-Surugadai, Chiyoda-ku, Tokyo 101-0062, Japan
- Department of Life Science and Applied Chemistry, Nagoya Institute of Technology, Gokiso-cho, Showa-ku, Nagoya 466-8555, Japan
- PRESTO, Japan Science and Technology Agency (JST), 4-1-8 Honcho, Kawaguchi, Saitama 332-0012, Japan
- Department of Nanopharmaceutical Sciences, Nagoya Institute of Technology, Gokiso-cho, Showa-ku, Nagoya 466-8555, Japan






















