
News & Events
News & Events
News
February 17, 2023
Pathogenesis of skeletal muscle lesions in spinal muscular atrophy
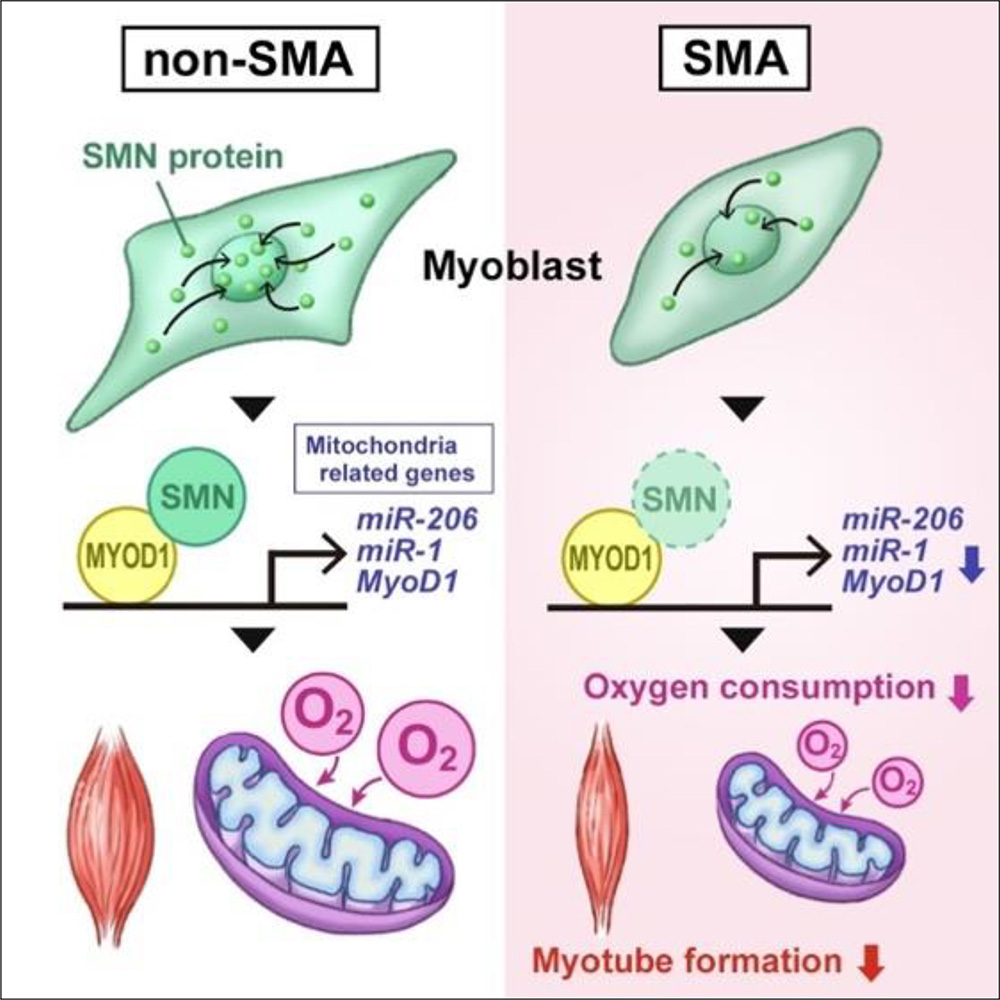
Spinal Muscular Atrophy (SMA) is a congenital neuromuscular disease caused by loss-of-function mutations in the SMN1 gene encoding the survival motor neuron (SMN) protein. Although SMA has traditionally been considered a motor neuron disease, skeletal muscle-specific lesions have also been recently reported, making skeletal muscle a new potential therapeutic target for SMA. In this study, the research group investigated the pathogenesis of SMA skeletal muscle lesions using myoblasts established from SMA patients-derived iPS (SMA iPS) cells.
The group also generated SMN1-deficient iPS cells by knocking down SMN1 in iPS cells derived from healthy individuals (non-SMA iPS cells) using shRNA. The research team found that both skeletal muscle cells differentiated from SMN1-deficient iPS cells and SMA iPS cells exhibited decreased mitochondrial oxygen consumption. Furthermore, the expression levels of endogenous MYOD1, miR-1, and miR-206, critical to mitochondrial functions during skeletal muscle differentiation, were also reduced.
Next, the group investigated the function of SMN during skeletal muscle differentiation. Skeletal muscle differentiation from non-SMA iPS cells increased SMN1 expression after three days, and SMN proteins were distributed uniformly in the cell nuclei. These observations suggest that SMN interacted with DNA to control transcription during differentiation. Consistently, ChIP-qPCR revealed that SMN bound to the genomic region upstream of the MYOD1 transcription start site. Furthermore, analysis by co-immunoprecipitation confirmed that SMN and MYOD1 proteins physically interacted.
Previous studies have shown the transcription of miR-1 and miR-206 to be regulated by the binding of MYOD1 to their transcriptional regulatory regions. Therefore, the researchers examined the binding of SMN to the transcriptional regulatory sites of miR-1 and miR-206 by ChIP-qPCR and found that it was also bound to these miRNA genes. These results suggest that SMN is specifically involved in the transcriptional regulation of MYOD1, miR-1, and miR-206 during skeletal muscle differentiation.
Finally, the group examined whether introducing miR-1 and miR-206, downregulated in the SMA model, into skeletal muscle would be an effective treatment for SMA. The results showed increased mitochondrial oxygen consumption and improved myotube cell formation following the introduction of miR-1 and miR-206. Functionally, miR-1 and miR-206 transfections restored the contraction velocity of myotube cells from SMA model mice to levels comparable to those of wild-type myotube cells. These results suggest that miR-1 and miR-206 are potent targets for rescuing the skeletal muscle pathology in SMA.
This study has identified one of the pathogenic mechanisms of skeletal muscle lesions in SMA. The research team hopes their work will lead to the development of novel therapies by targeting skeletal muscles in the future.
The results of this research were published online in Life Science Alliance on January 5, 2023.
Paper Details
- Journal: Life Science Alliance
- Title: SMN promotes mitochondrial metabolic maturation during maturation by regulating the MYOD-miRNA axis
- Authors:
Akihiro Ikenaka1, Yohko Kitagawa1, Michiko Yoshida2, Chuang-Yu, Lin1,3, Akira Niwa1,
Tatsutoshi Nakahata4, Megumu K. Saito1*
* Corresponding Author - Author Affiliations:
- Department of Clinical Application, Center for iPS Cell Research and Application, Kyoto University, Kyoto, Japan
- Department of Pediatrics, Kyoto Prefectural University of Medicine, Kyoto, Japan
- Department of Biomedical Science and Environmental Biology, Kaohsiung Medical University, Kaohsiung, Taiwan
- Drug Discovery Technology Development Office, Center for iPS Cell Research and Application, Kyoto University, Kyoto, Japan






















