
Research Activities
Research Activities
Publications
August 09, 2017
Bacteria changes mammalian cell identity
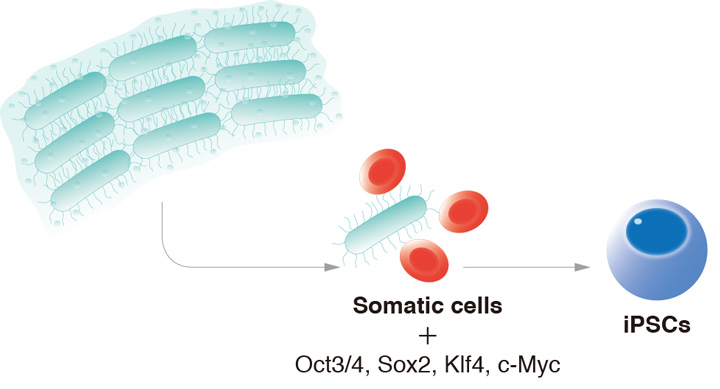
CiRA researchers show bacterial proteins enhance the reprogramming of mouse cells to iPS cells.
The discovery of iPS cells demonstrated that a cell's identity could be change through the activation and repression of certain genes. While it is now possible to reprogram cells from one identity to another, the low efficiency limits the number of experiments and medical applications. A new study by CiRA researchers reports that using proteins from bacteria can enhance the reprogramming efficiency, giving light to how xenoproteins can regulate cell behavior.
CiRA Junior Associate Professor Shinji Masui, who led the study, notes that a number of chemical compounds have been found to improve reprogramming efficiency, but efficiency remains dissatisfactory, and finding new ones is not trivial.
"Several small molecules are known to enhance reprogramming, but the search is laborious. Bacterial proteins are easier to screen and prepare," he said.
According to Masui, scientists must screen tens of thousands of small molecules to find one candidate, whereas they need only screen tens of bacterial proteins.
Cell reprogramming to iPS cells was first demonstrated by transducing into mouse cells the four Yamanaka factors. In the new study, Masui and his colleagues tested 30 genes from the bacterium Wolbachia pipientis and found that introducing the protein coded from one of these genes with the Yamanaka factors into mouse cells changed the reprogramming efficiency. However, the effect depended on the cell type, as both mouse fibroblasts and mouse neural progenitor cells could be reprogrammed more efficiently, but mouse hepatoblasts became more resistant to reprogramming when the bacterial protein was added.
Masui admits that the dependency on cell deserves more study. Further analysis showed that the protein enhanced reprogramming by repressing genes that sustain cell identity, suggesting perhaps it may have to do with how xenoproteins interact with specific genes in the cell that accelerate the reprogramming.
"Wolbachia pipientis proteins downregulated cytoskeletal genes. Cell morphology is very important for cell identity. It determines how the cell interacts with its environment," said Masui.
The discovery that xenoproteins bind to cytoskeletal proteins suggests other methods that suppress these proteins could also benefit cell reprogramming science.
Masui is equally excited about the implications of bacteria influencing cell identity.
"Infections and diseases change cell behavior. The cell changes its identity. Understanding bacterial and mammalian cell interactions will allow us to understand disease development," he said.
Paper Details
- Journal: Genes to Cells
- Title: Artificial acceleration of mammalian cell reprogramming by bacterial proteins
- Authors: Ikeda T1, Uchiyama I2, Iwasaki M1, Sasaki T3, Nakagawa M1, Okita K1, and Masui S1
- Author Affiliations:
- Center for iPS Cell Research and Application (CiRA), Kyoto University, Kyoto, Japan
- National Institute for Basic Biology, National Institutes of Natural Sciences, Okazaki, Japan
- Honeybee Science Research Center, Tamagawa University Research Institute, Tokyo, Japan






















