
Research Activities
Research Activities
Publications
May 19, 2017
Synthetic gene circuits that turn gene editing on and off
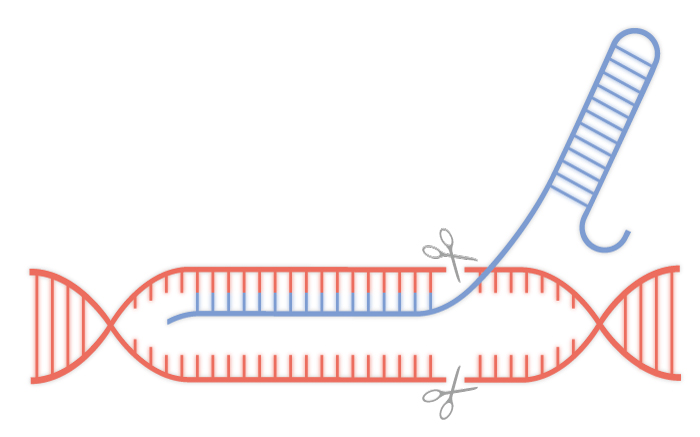
New RNA technology regulates the CRISPR-Cas9 system to
genome edit target cells from a mixed population.
used to regulate the CRISPR-Cas9 system inside cells.
The discovery of the CRISPR-Cas9 system less than a decade ago has made gene editing simple and precise. Once inside the cell, the Cas9 protein is guided into the nucleus where it edits the target gene. While powerful, scientists struggle to assure that the system will only edit the target cells of a mixed population. The Hirohide Saito lab now reports new synthetic biotechnology that achieves this goal.
The new biotechnology tool, miR-Cas9 switch, is a variation of the miRNA switch, which Saito first reported two years ago and describes as, "genetic circuits that control gene expression in response to miRNA. We can design the switch that responds to miRNA exclusive to a cell type, so that the gene is expressed or not expressed in only that type. "
Saito's miRNA switches are composed of a target site that binds to a specific miRNA sequence and RNA that codes for a specific protein, such as Cas9. "We can change the target site and the RNA sequence arbitrarily and independently," he said.
Knowing this, Moe Hirosawa, a graduate student in the Saito lab, investigated whether "we can control gene editing in different cells by making the expression of Cas9 dependent on our miRNA switch."
Hirosawa prepared a Cas9 protein that edits the green fluorescent protein (GFP) gene and a miRNA switch that controlled Cas9 expression. Cells that did not express Cas9 protein turned green, while those that expressed Cas9 did not. To regulate the Cas9 expression, he set the target site to one that binds miR-302a, a type of miRNA that is found at extraordinarily higher levels in human iPS cells than other cell types.
Transfecting this miR-Cas9 switch into a mixture of iPS cells and other cells, he found only the former fluoresced green, demonstrating that miR-302a binding prevented Cas9 synthesis in iPS cells and more importantly that he could program the CRISPR-Cas9 system to function in only specific cell types. Changing the target site to one that binds to miR-21, a miRNA highly expressed in HeLa cells, but not iPS cells, he used his miR-Cas9 switch to fluoresce HeLa cells but not iPS cells.
However, in this current design, the miR-Cas9 switch only edits a gene if miRNA does not bind to the target site, making it an OFF switch. To create an ON miR-Cas9 switch further modifications were needed.
"We added a RNA-binding protein component to the switch. This turned the switch from OFF to ON when miRNA binds to the mRNA," said Saito.
The preferred RNA-binding protein in the Saito lab is L7Ae. In the modified miR-Cas9 switch, Cas9 expression depends on the presence of L7Ae and L7Ae expression depends on the binding of miRNA to the target site.
"It is a little more complicated, because we have to prepare two synthetic mRNAs," explains Hirosawa. "One for L7Ae and one for Cas9."
The team believes that though their technology is preliminary, it could provide an effective way to selectively control gene editing and cell-fate among heterogeneous cell populations.
Paper Details
- Journal: Nucleic Acids Research
- Title: Cell-type-specific genome editing with a microRNA-responsive CRISPR-Cas9 switch
- Authors: Hirosawa M1,2, Fujita Y1, Parr CJC1, Hayashi K1, Kashida S1, Hotta A1, Woltjen K1,3, and Saito H1
- Author Affiliations:
- Center for iPS Cell Research and Application (CiRA), Kyoto University, Kyoto, Japan
- Graduate School of Medicine, Kyoto University, Kyoto, Japan
- Hakubi Center for Advanced Research, Kyoto University, Kyoto, Japan






















