
Research Activities
Research Activities
Publications
February 28, 2020
Stem cells are not afraid of water
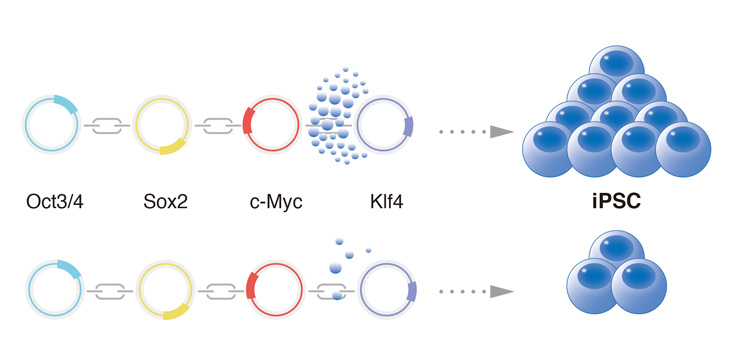
The hydrophobicity of the linker for KLF4 influences iPS cell generation and stability.
Water molecules around KLF4 indicate the hydrophobicity.
A new study in Stem Cell Reports by CiRA scientists identifies a seemingly simple technical detail that influences the probability of reprogramming cells into iPS cells. The study reports that a protein linker commonly used to connect reprogramming factors affects the stability of one factor in particular, KLF4. Modifying the hydrophobicity of the protein linker enhanced KLF4 stability and thus enhanced the reprogramming efficiency. Overall, the study shows that besides understanding the biology of cell programming, understanding the enabling biotechnology could have profound effects on iPS cell production.
The original iPS cells were made by inducing the transient expression of four genes, or Yamanaka factors, including KLF4. To initiate the reprogramming process, scientists inserted four "monocistronic" gene cassettes, each carrying one Yamanaka factor, into the cell.
However, explains CiRA Associate Professor Knut Woltjen, the monocistronic approach is challenging.
"Getting all four genes into one cell is not trivial. Much easier is to make one cassette that turns on all four genes," he says.
Thus, combining the factors' genes into one "polycistronic" cassette has become a preferred method. In these polycistronic cassettes, the four genes are linked together with a series of 2A peptide sequences, a biotechnology trick borrowed from viruses that compact their genomes.
The Woltjen laboratory has published several papers showing that the structure of KLF4 in polycistronic cassettes can have a major effect on the reprogramming efficiency. In particular, they found that of two KLF4 isoforms, the longer version generated more protein and was more efficient at cell reprogramming. However, their results did not point to any clear reason.
"We knew that the amount of KLF4 protein affected many of the reprogramming phenotypes, including the mesenchymal-to-epithelial transition and partial reprogramming. What we did not know was the underlying cause," says Dr. Harunobu Kagawa of the Institute of Molecular Biotechology (IMBA) in Vienna, who had written a previous study about the isoforms with Woltjen and is a coauthor on this one.
Although the 2A peptide approach is standard for the preparation of polycistronic cassettes, some evidence suggests that it can disrupt the stability of the proteins expressed by the linked genes.
"That's why we speculated the effect was due to the 2A peptide," says Dr. Anika Reinhardt a post-doctoral fellow who was funded by the iPS Cell Research Fund and has since moved to a company in Berlin, Germany, as a scientist.
The researchers went on to compare several different 2A peptide sequences and identified the inclusion of the amino acid glutamate as maximizing KLF4 expression. This was attributed to glutamate reducing the protein hydrophobicity, thus stabilizing KLF4. The lower hydrophobicity was found to reduce susceptibility to proteasome activity, which degrades proteins.
"Using glutamate, we were able to design our cassette so that the shorter KLF4 isoform could reprogram cells as effectively as the longer KLF4 isoform," remarks Reinhardt.
Beyond the implications to cell reprogramming, Woltjen stresses that the study shows the importance of carefully considering how artificial DNA constructs can modify gene expressions.
"Our results suggest that a single amino acid can be used to counteract the instability of susceptible proteins. It's remarkable how seemingly minor modifications can achieve such a major effect."
Paper Details
- Journal: Stem Cell Reports
- Title: N-terminal amino acids determine KLF4 protein stability in 2A peptide-linked polycistronic reprogramming constructs
- Authors: Anika Reinhardt, Harunobu Kagawa and Knut Woltjen
- Author Affiliations: Center for iPS Cell Research and Application, Kyoto University, Kyoto, Japan






















